Velociraptor 0.6.7 Release
I am very excited to announce the latest Velociraptor release 0.6.7 is now out. This release has been in the making for a few months now and has a lot of new features and bug fixes.
In this post I will discuss some of the interesting new features.
NTFS Parser changes
In this release the NTFS parser was improved significantly. The main areas of developments were around better support for NTFS compressed and sparse files and better path reconstruction.
In NTFS there is a Master File Table (MFT) containing a record for
each file on the filesystem. The MFT entry describes a file by
attaching several attributes to the file. Some of these attributes are
$FILE_NAME attributes representing the file names of the file.
In NTFS a file may have multiple names. Normally, files have a long
file name and a short filename. Each $FILE_NAME record also contains
a reference to the parent MFT entry of its directory.
When Velociraptor parses the MFT it attempts to reconstruct the full
path of each entry by traversing the parent MFT entry, recovering its
name etc. Previously, Velociraptor used one of the $FILE_NAME
records (usually the long file name) to determine the parent MFT
entry. However, this is not strictly correct as each $FILE_NAME
record can a different parent directory. This surprising property
of NTFS is called hard links.
You can play with this property using the fsutil program. The
following adds a hard link to the program at
C:/users/test/downloads/X.txt into a different directory.
C:> fsutil hardlink create c:\Users\Administrator\Y.txt c:\Users\Administrator\downloads\X.txt
Hardlink created for c:\Users\Administrator\Y.txt <<===>> c:\Users\Administrator\downloads\X.txt
The same file in NTFS can exist in multiple directories at the same
time by use of hard links. The filesystem simply adds a new
$FILE_NAME entry to the MFT entry for the file pointing at another
parent directory MFT entry.
Therefore, when scanning the MFT, Velociraptor needs to report all possible directories each MFT entry can exist in (There can be many such directories, since each directory can have hard links itself too).
As a rule an MFT Entry can represent many files in different directories!
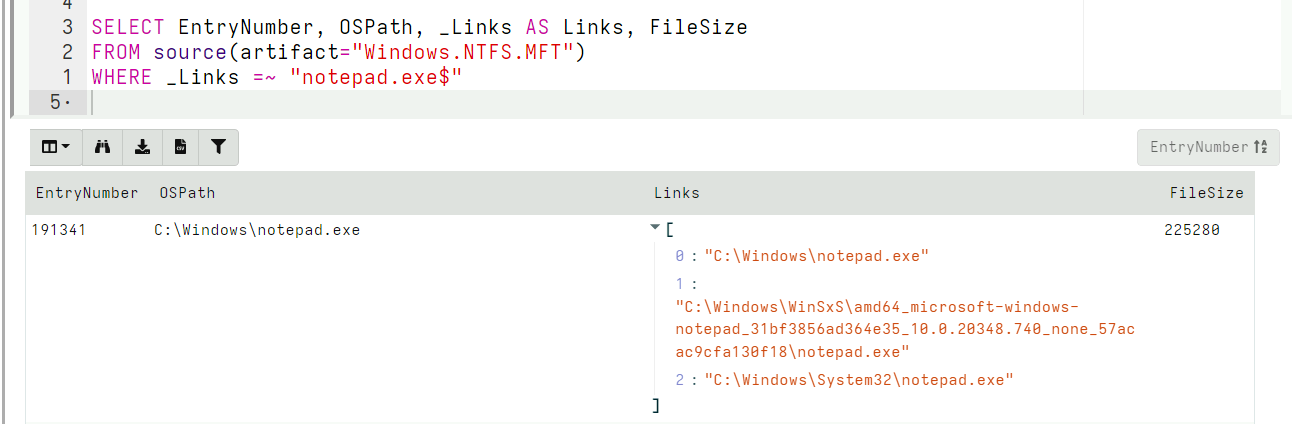
Reassembling paths from MFT entries
When Velociraptor attempts to reassemble the path from an unallocated MFT entry it might encounter an error where the parent MFT entry indicated has already been used for some other file or directory.
In previous versions, Velociraptor simply reported these parents as potential parts of the full path, since for unallocated entries the path reconstruction is best effort. This lead to confusion among users with often nonsensical paths reported for unallocated entries.
In the latest release, Velociraptor is more strict in reporting parents of unallocated MFT entries, also ensuring that the MFT sequence numbers match. If the parent’s MFT entry sequence number does not match, Velociraptor’s path reconstruction indicates this as an error path.
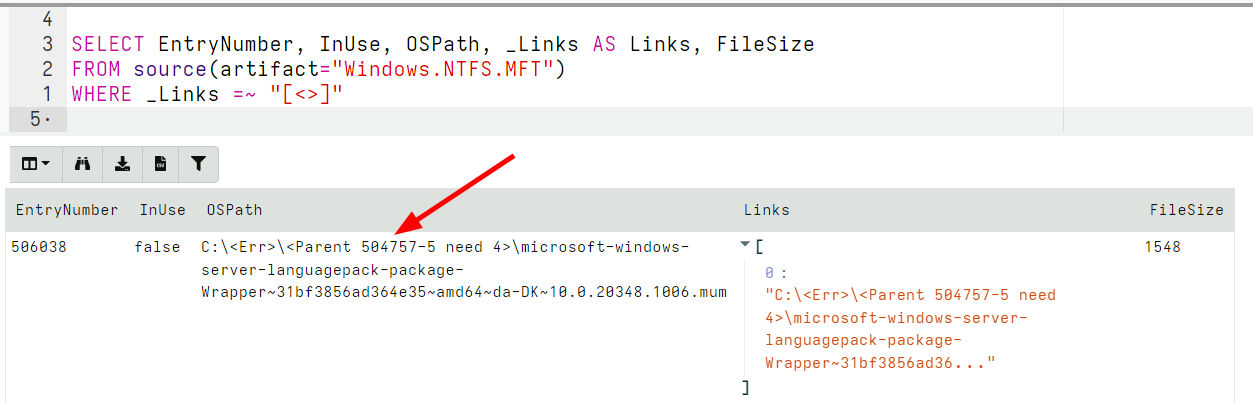
In the above example, the parent’s MFT entry has a sequence number of 5, but we need a sequence number of 4 to match it. Therefore the parent’s MFT entry is rejected and instead we report the error as the path.
The offline collection and encryption
Velociraptor’s offline collector is a pre-configured Velociraptor binary which is designed to be a single shot acquisition tool. You can build an Offline Collector by following the documentation. The Offline Collector does not require access to the server, instead simply collecting the specified artifacts into a Zip file (which can subsequently be uploaded to the cloud, or simply shared with the DFIR experts for further analysis).
Previously, Velociraptor only supported encrypting the Zip archive using a password. This is problematic because the password had to be embedded inside the collector configuration and so could be viewed by anyone with access to the binary.
In the latest release, Velociraptor supports asymmetric encryption to
protect the acquisition Zip file. There are two asymmetric schemes:
X509 encryption and PGP encryption. Having asymmetric encryption
improves security greatly because only the public key needs to be
included in the collector configuration. Dumping the configuration
from the collection is not sufficient to be able to decrypt the
collected data - the corresponding private key is also required!
This is extremely important for forensic collections since these will often contain sensitive and PII information.
Using this new feature is also extremely easy: One simply selects the X509 encryption scheme during the configuration of the offline collector in the GUI.
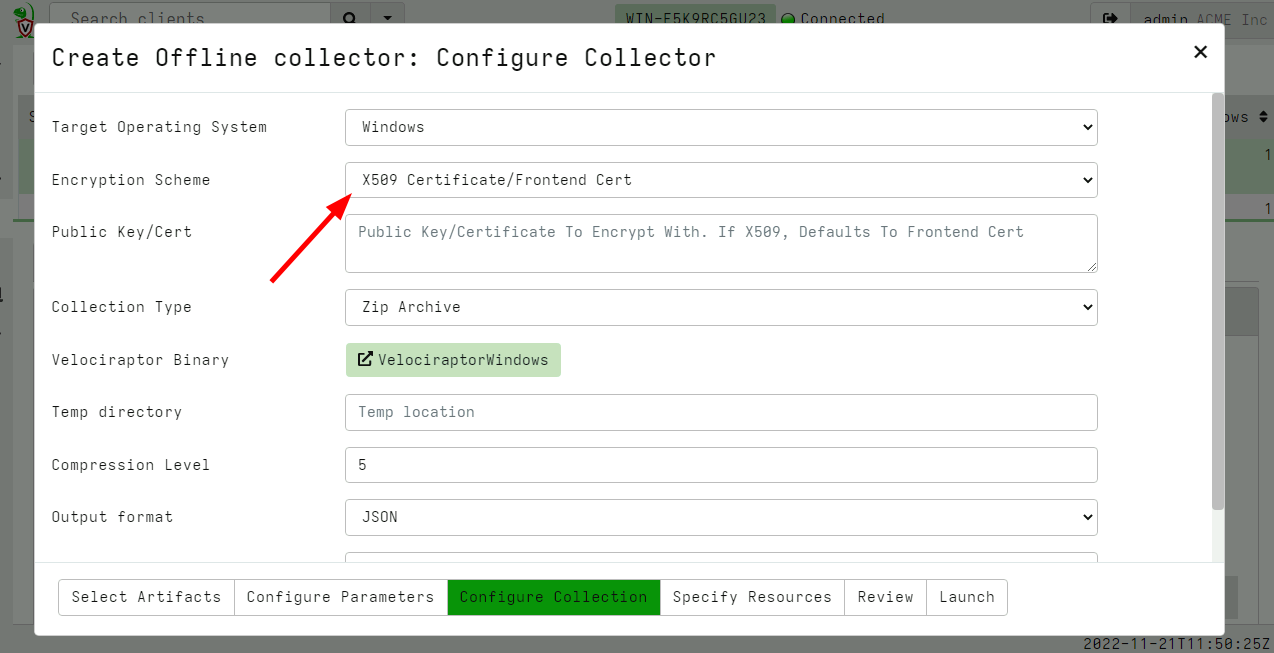
You can specify any X509 certificate here, but if you do not specify any, Velociraptor will use the server’s X509 certificate instead.
Velociraptor will generate a random password to encrypt the Zip file with, and then encrypt this password using the X509 certificate.
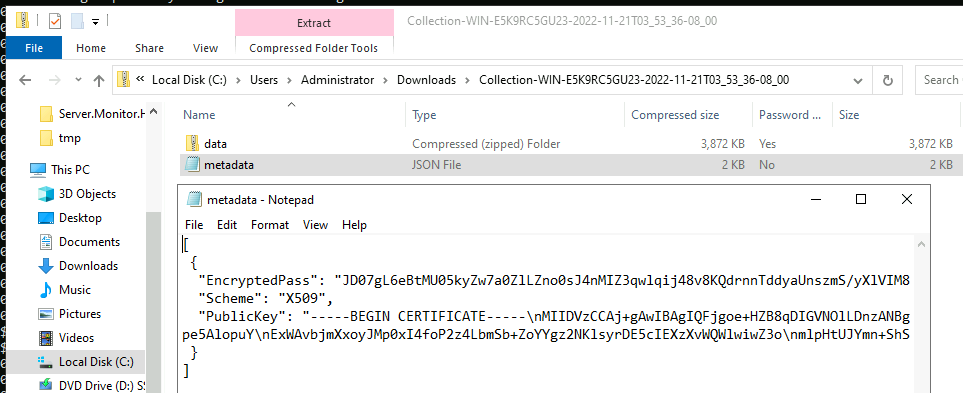
Since the ZIP standard does not encrypt the file names, Velociraptor
embed a second zip called data.zip inside the container. The above
illustrated the encrypted data zip file and the metadata file that
describes the encrypted password.
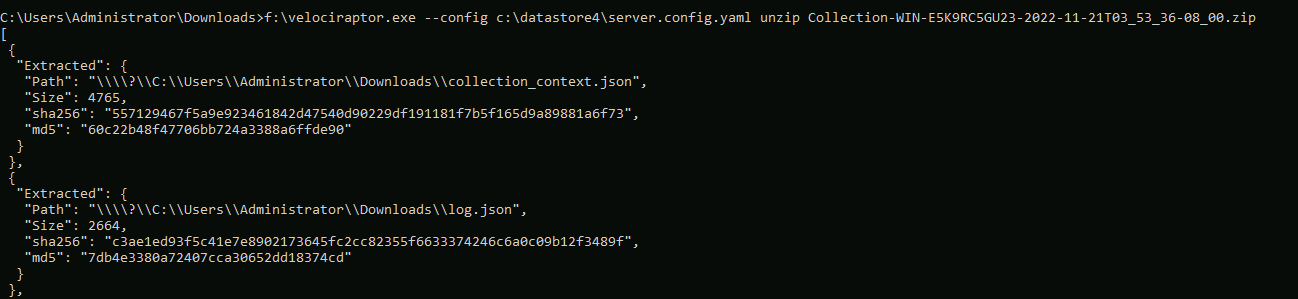
Because the password used to encrypt the container is not known and
needs to be derived from the X509 private key, we must use
Velociraptor itself to decrypt the container (i.e. we can not use
e.g. 7zip).
Importing offline collections
Originally the offline collector feature was designed as a way to collect the exact same VQL artifacts that Velociraptor allows in the usual client-server model in situations where installing the Velociraptor client was not possible. The same artifacts can be collected into a zip file.
As Velociraptor’s post processing capabilities improved (using notebooks and server side VQL to enrich the analysis), people naturally wanted to use Velociraptor to post process offline collections too.
Previously Velociraptor did have the Server.Utils.ImportCollection
artifact to allow an offline collection to be imported into
Velociraptor but this did not work well because the offline collector
simply did not include enough information in the Zip file to
sufficiently emulate the GUI’s collection views.
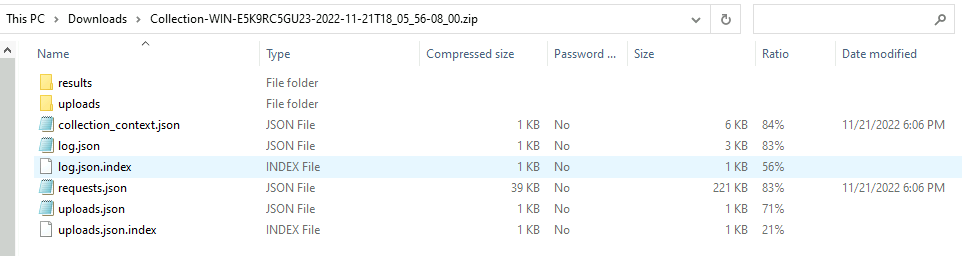
In the recent release, the offline collector was updated to add more detailed information to the collection zip, allowing it to be easily imported.
Exporting and Importing collections
Velociraptor has previously had the ability to export collections and hunts from the GUI directly, mainly so they can be processed by external tools.
But there was no way to import those collections back into the GUI. We just never imagined this would be a useful feature!
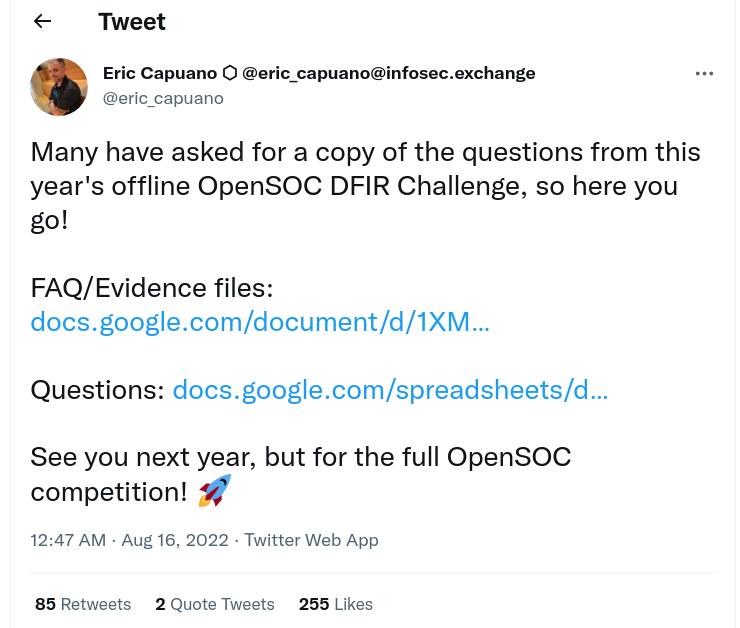
Recently Eric Capuano from ReconInfosec shared some data from an exercise using Velociraptor and people wanted to import this into their own Velociraptor installations so they can run notebook post processing on the data themselves.
Our community has spoken though! This is a useful feature!
In the latest release exported files from the GUI use the same container format as the offline collector and therefore can be imported into a different Velociraptor installation seamlessly.
Handling of sparse files
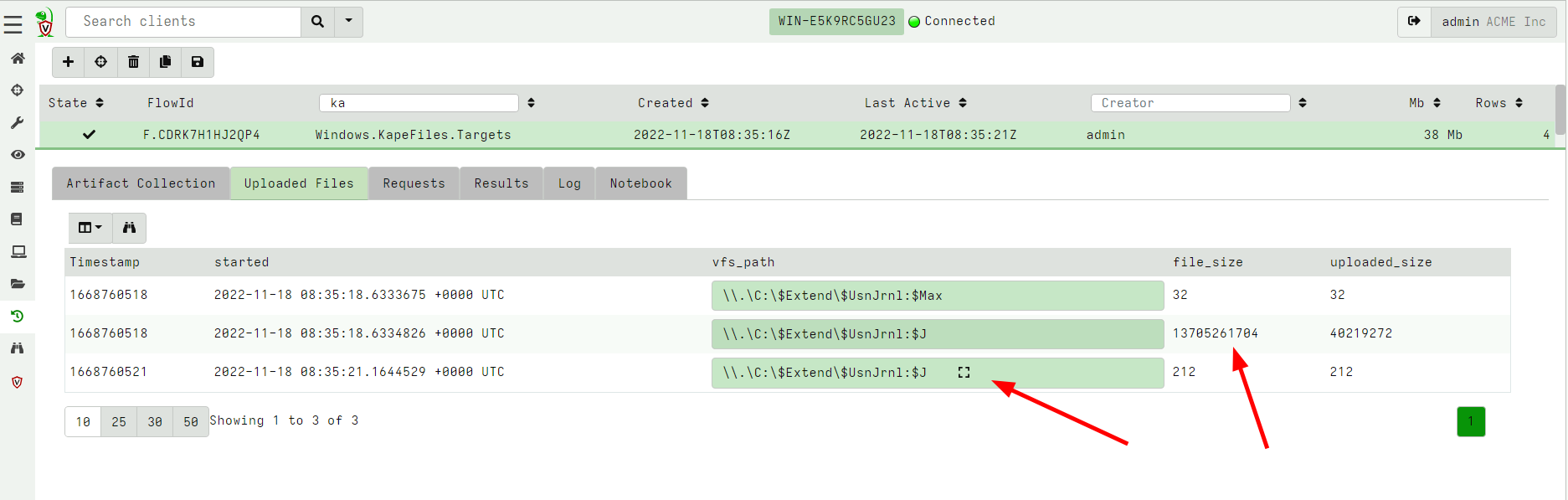
When collecting files from the endpoint using the NTFS accessor we quite often encounter sparse files. These are files with large unallocated holes in them. The most extreme sparse file is the USN Journal.
In the above example the USN journal size is reported to be 1.3Gb but in reality only about 40mb is occupied on disk. When collecting this file, Velociraptor only collects the real data and marks the file as sparse. The Zip file will contains an index file which specifies how to reassemble the file into its original form.
While Velociraptor stores the file internally in an efficient way, when exporting the file for use by other tools, they might expect the file to be properly padded out (so that file offsets are correct).
Velociraptor now allows the user the choice of exporting an individual file in a padded form (with sparse regions padded). This can also be applied to the entire Zip export in the GUI.
For very large sparse files, it makes no sense to pad so much data out (Some USN journal files are in the TB region), so Velociraptor implements a limit on padding of very sparse files.
Parsing User Registry Hives
Many Velociraptor artifacts simply parse keys and values from the registry to detect indicators. Velociraptor offers two methods of accessing the registry:
- Using the Windows APIs
- Employing the built in raw registry parser to parse the hive files.
While the first method is very intuitive and easy to use, it is often problematic. Using the APIs requires the user hive to be mounted. Normally the user hive is only mounted when a user logs in. Therefore querying registry keys in the user hive will only work on users that are currently logged in at the time of the check and miss other users (which are not currently logged in so their hive is not mounted).
To illustrate this problem consider the
Windows.Registry.Sysinternals.Eulacheck artifact which checks the
keys in HKEY_USERS\*\Software\Sysinternals\* for the Sysinternals
EULA value.
In previous versions of Velociraptor, this artifact simply used the windows API to check these keys/values and completely missed any users that were not logged in.
While this issue is know, previously users had to employ complex VQL
to customize the query so it can search the raw NTUSER.DAT files in
each user registry. This is more difficult to maintain since it
requires two separate types of artifact for the same indicator.
With the advent of Velociraptor’s dead disk capabilities it is
possible to run a VQL query in a “virtualized” context consisting of a
remapped environment. The end result is that the same VQL query can be
used to run on raw registry hives. It is now trivial to apply the same
generic registry artifact to a raw registry parse.
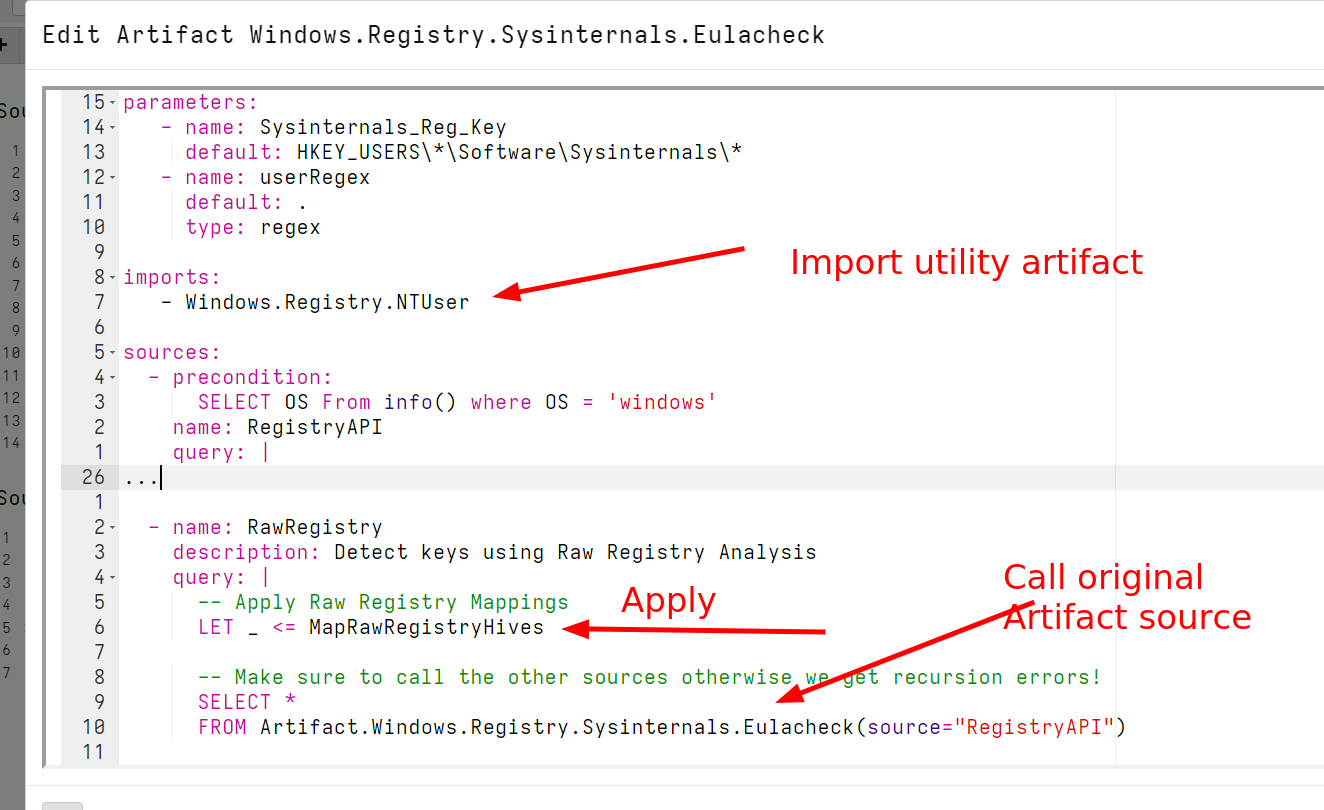
All that is required to add raw registry capabilities to any registry artifact is:
- Import the
Windows.Registry.NTUserartifact - Use the
MapRawRegistryHiveshelper function from that artifact to set up the mappings automatically. - Call the original registry query using the
registryaccessor. In the background this will be remapped to the raw registry accessor automatically.
Conclusions
There are many more new features and bug fixes in the latest release.
If you like the new features, take Velociraptor for a spin! It is a available on GitHub under an open source license. As always please file issues on the bug tracker or ask questions on our mailing list velociraptor-discuss@googlegroups.com . You can also chat with us directly on Discord. .